GIFT Shuangjia Zheng's Team Publishes Cell Systems Cover Article: COSMOS Tackles the Critical Challenge of Precise Protein Function Annotation
Protein function annotation is not only a cornerstone for understanding life mechanisms, discovering drug targets, and advancing biotechnological applications, but also a vital bridge linking genomic information to biological activities. However, the traditional Gene Ontology (GO) framework remains heavily reliant on labor-intensive and costly wet-lab experiments, leaving the majority of protein functions underexplored. For example, over 80% of UniProt proteins lack curated functional annotations. Furthermore, hundreds of new functional categories are introduced each year, posing severe zero-shot and few-shot challenges for predictive models. The reliance of the existing approaches on sequence similarity or structural features limits their applicability to low-homology proteins or underexplored classes. Therefore, accurate prediction from sparse, dynamic functional annotation data has become a critical bottleneck in computational biology.


To address these challenges, a research team led by Assistant Professor Shuangjia Zheng at the Global Institute of Future Technology, Shanghai Jiao Tong University, introduced COSMOS, a foundation model that constructs a knowledge graph integrating protein interactions, structural similarity and hierarchical functional relationships. By leveraging an inductive subgraph learning framework, COSMOS achieves accurate, interpretable prediction of protein functions, including for unknown, sparsely annotated, and low-homology proteins, and offers a new paradigm for protein function annotation. The paper titled "Context-informed subgraph foundation models enable interpretable protein-function prediction" was published as the cover article on Cell Systems.
The COSMOS model innovatively integrates subgraph deep learning with a protein-GO knowledge graph (KG). First, the researchers construct an enriched multi-source protein-function KG containing nearly 8 million functional semantic relationships by integrating protein function annotations, GO hierarchy, PPIs, and structural proteomics data. Instead of conventional global graph embedding methods, COSMOS then extracts local subgraphs specific to each candidate protein–function pair and employs a multi-relational message-passing neural network to precisely capture the logical topological structures embedded within these subgraphs. The model is pre-trained using a noise-contrastive loss and ultimately outputs an association score between a protein and a function.

Figure 1: Illustration of the COSMOS framework and its application
In multiple challenging experimental settings, COSMOS demonstrated superior performance over existing methods. In zero-shot prediction tasks where training and testing GO categories share no overlap, COSMOS achieved an Fmax of 0.915 for molecular functions, a 21.8% improvement over DeepGOGATSE. In few-shot and low-homology classes, COSMOS also displayed robust performance, outperforming the best baseline models by 13.8% and 19.9% in Fmax, respectively. Ablation studies confirmed that integrating protein interaction and structural similarity information accounted for over 90% of the model's performance, highlighting the critical role of multi-source data fusion. Embedding space analysis revealed that COSMOS can clearly differentiate correct protein–function associations from incorrect ones, with protein–function pairs sharing the same functional categories naturally clustering in the embedding space, further validating the model's capacity to extract topological features that effectively reflect functional associations.
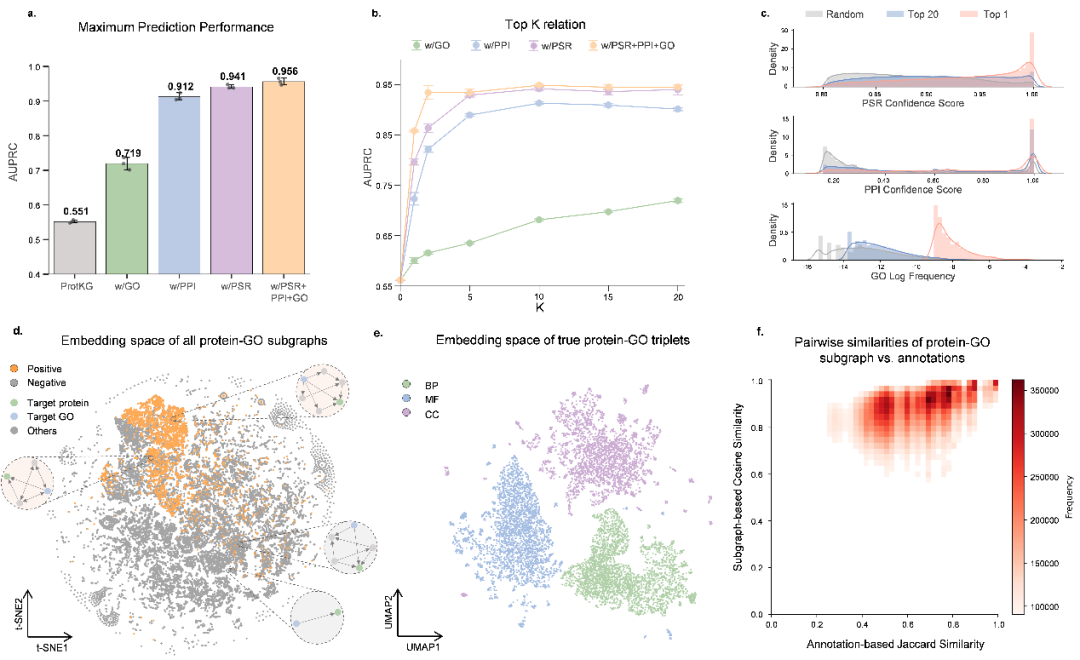
Figure 2: Performance analysis of COSMOS with different proposed techniques
Beyond theoretical benchmarks, COSMOS demonstrated substantial value in real-world scenarios. A case study on the protein SIRT6 showed that COSMOS not only predicted its classical histone deacetylase activity but also identified its defatty-acylase activity toward long-chain fatty acyl lysines - a function discovered only after the model's training data cutoff date - and provided a clear subgraph pathway as predictive evidence. Furthermore, cross-species testing across humans, mice, and yeast consistently showed COSMOS's leading performance, with a 12% improvement in average AUPRC over the best baseline on mice datasets, demonstrating its strong generalizability.
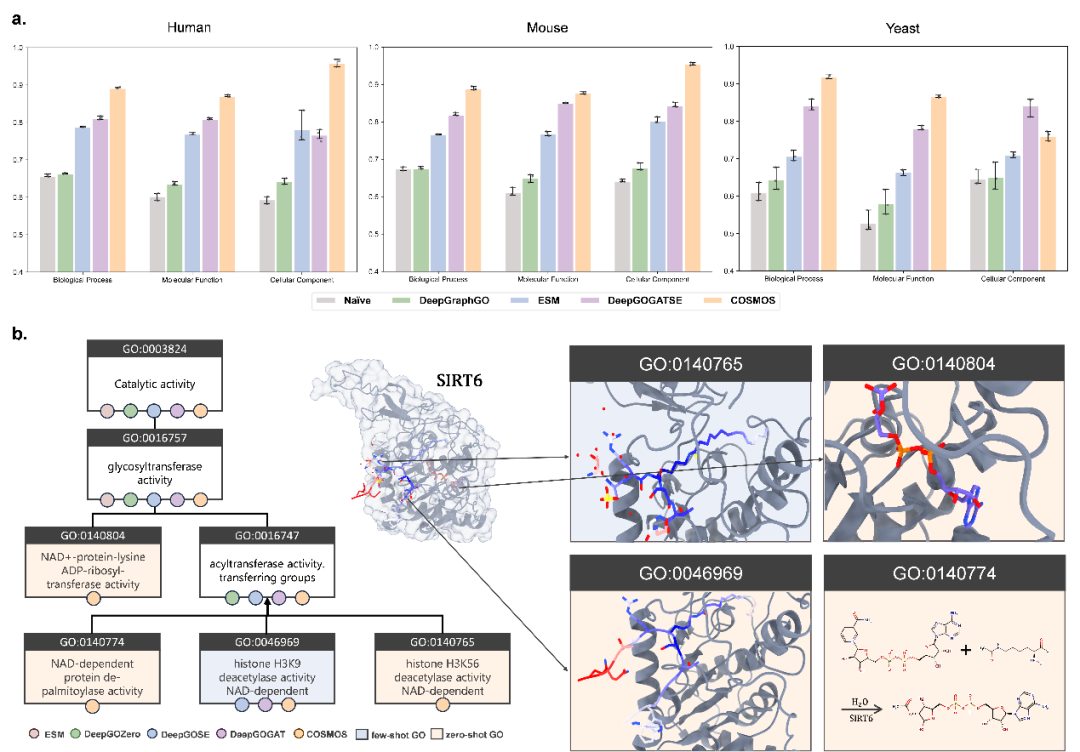
Figure 3: Performance of COSMOS AUPRC and validation examples
In summary, COSMOS establishes a powerful framework that integrates structural information, network interactions and functional semantics for protein function prediction, breaking free of dependence on sequence homology and demonstrating significant advantages in complex scenarios such as zero-shot, few-shot, and low-homology settings. COSMOS provides an orthogonal and complementary approach to existing protein-function prediction methods, further improving predictive performance. The team has open-sourced both the model and data, providing the research community with a highly interpretable and generalizable protein function annotation tool. COSMOS is expected to shorten the annotation cycle for novel proteins and accelerate advances in functional genomics, drug target discovery, and synthetic biology.
Zhuomin Zhou, a master's student of SJTU, is the first author of the paper. Assistant Professor Shuangjia Zheng serves as the sole corresponding author.
Paper Title:
Context-informed subgraph foundation models enable interpretable protein-function prediction
Paper Link:
https://www.cell.com/cell-systems/fulltext/S2405-4712(26)00017-7




