GIFT Shuangjia Zheng's Team Publishes in Nature Catalysis: Multimodal Foundation Model EnzymeCAGE Enables Accurate Enzyme Catalysis Prediction
On February 12, 2026, a research team led by Shuangjia Zheng at the Global Institute of Future Technology, SJTU, published a paper in Nature Catalysis entitled "A Geometric Foundation Model for Enzyme Retrieval with Evolutionary Insights." Conducted in collaboration with researchers from The Hong Kong University of Science and Technology, Massachusetts Institute of Technology, Sun Yat-sen University, and other international institutions, the study introduces EnzymeCAGE, a geometric foundation model that integrates protein structural information with evolutionary signals to achieve high-precision matching between biochemical reactions and enzymes, demonstrating significant advantages in the design and reconstruction of biosynthetic pathways, and providing a new technological pathway for AI-driven enzyme discovery and biomanufacturing.

Enzymes are nature's catalysts—highly efficient and specific "molecular machines" that drive complex biochemical transformations under mild conditions. However, their intricate structures and catalytic mechanisms have long posed significant challenges for functional annotation. This has left numerous biochemical reactions as "orphan reactions" lacking known enzymatic counterparts, creating a critical bottleneck in biosynthetic pathway elucidation and the development of synthetic biology. Existing methods mainly rely on sequence homology or functional classification and perform poorly in low-similarity scenarios. Therefore, overcoming sequence limitations and establishing a precise structure-function matching mechanism represents a crucial challenge.
To address this challenge, the researchers proposed EnzymeCAGE, a novel dual-drive structure combining geometric perception and evolutionary insights. By capturing three-dimensional features of enzyme catalytic pockets and leveraging protein language models to characterize both local structures and global sequences, the model provides a comprehensive representation of enzyme function. It further utilizes atomic mapping to analyze reaction center transformations from substrates to products, constructing precise reaction fingerprints. Ultimately, a geometrically enhanced interaction module quantifies the spatial compatibility between enzymes and reaction molecules, delivering catalytic compatibility scores that enable highly accurate enzyme recruitment and function prediction.
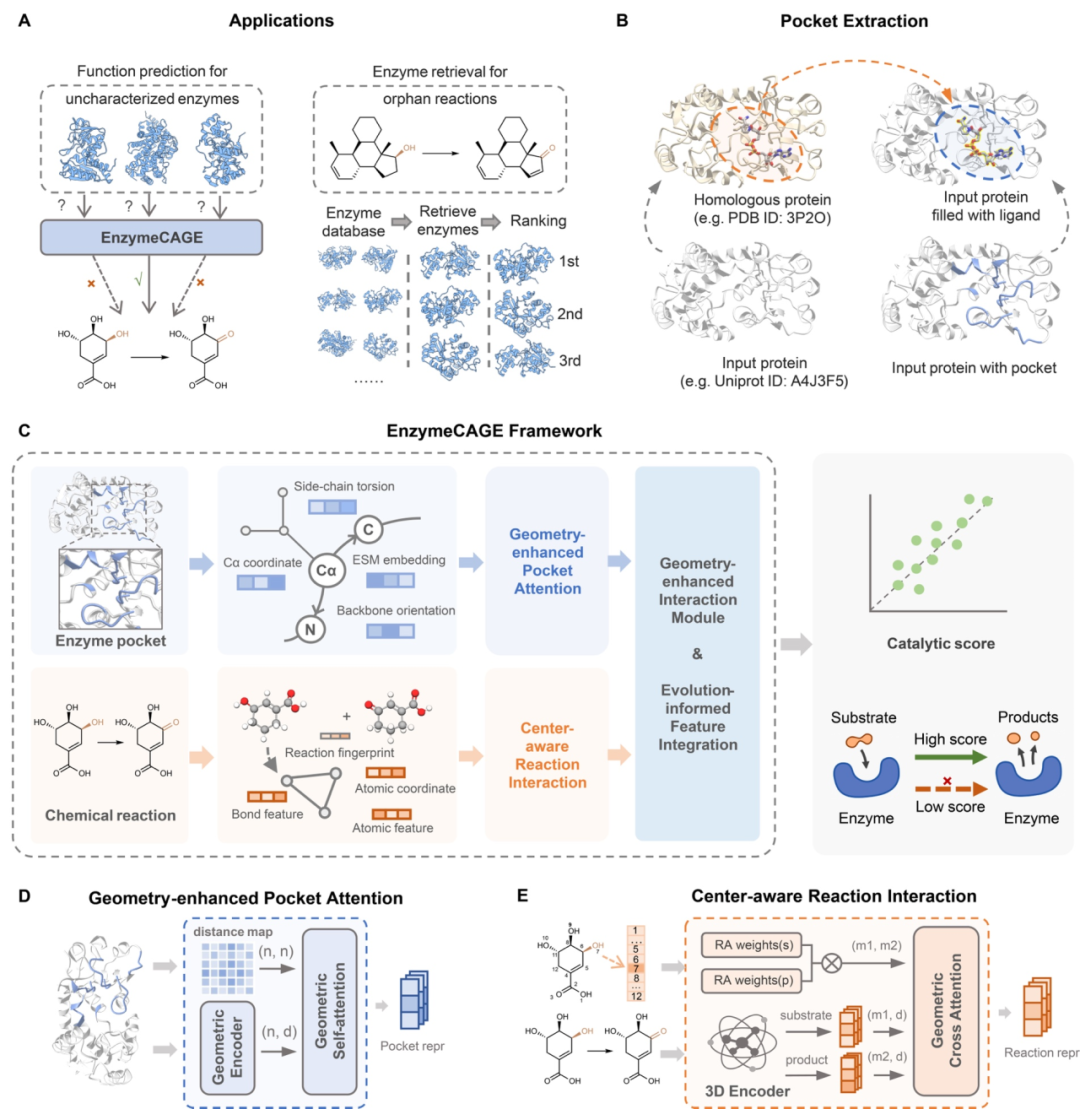
Figure 1: Overview of EnzymeCAGE
Extensive benchmarking demonstrates EnzymeCAGE's superiority over traditional methods. In evaluations involving hundreds of previously uncharacterized enzymes, the model achieved a top-10 success rate of 58%, outperforming tools such as MMseqs2 and CLIPZyme. Notably, EnzymeCAGE effectively overcomes sequence homology limitations, accurately identifying functional enzymes even in scenarios of extreme sequence divergence. In orphan reaction identification tasks, its efficiency surpasses conventional approaches by 41%, effectively filling gaps in metabolic networks. Moreover, through targeted optimization for key industrial enzyme families such as cytochromes P450, the model exhibits enhanced adaptability and consistently ranks specific targets among top predictions, demonstrating its broad applicability and practical utility.
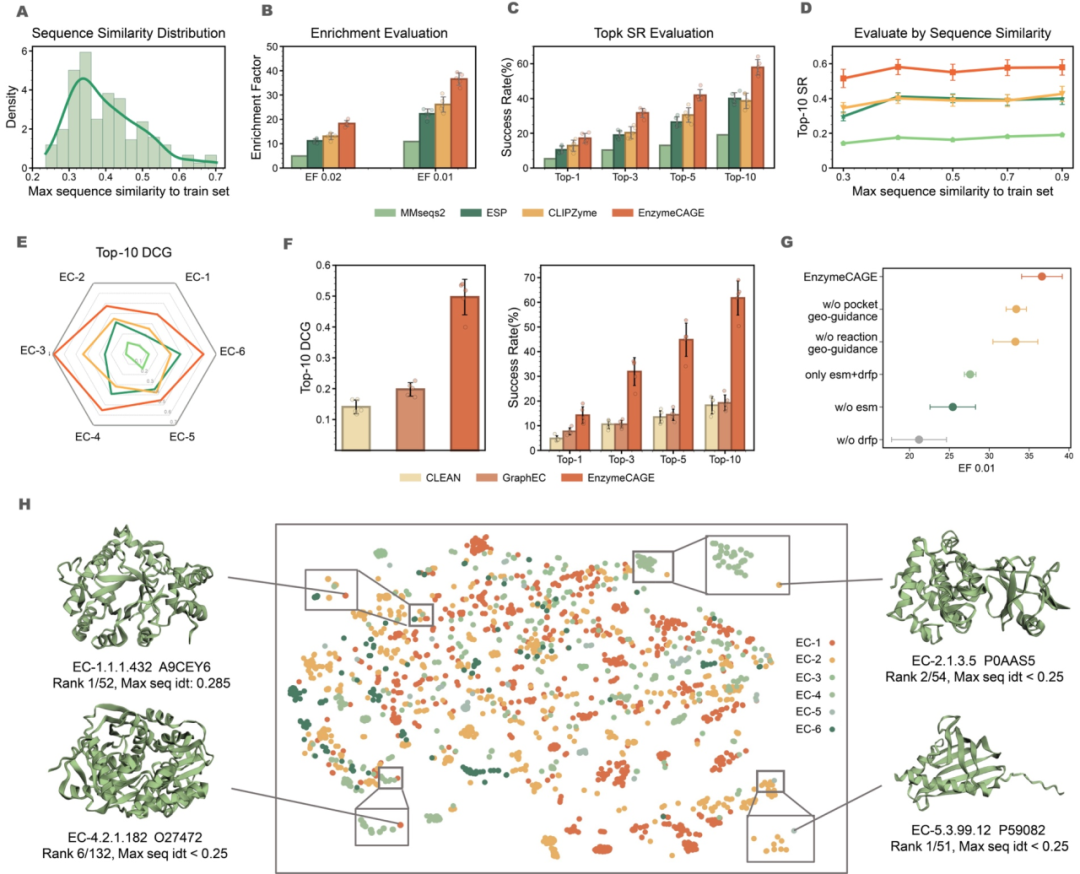
Figure 2: EnzymeCAGE's performance on enzyme recruitment and function prediction tests
Beyond theoretical advancements, EnzymeCAGE has shown substantial industrial value in real-world biomanufacturing applications. In the biosynthetic exploration of the anticancer compound withanolide, the model successfully identified three key P450 enzymes from hundreds of candidate sequences with highly favorable predictive rankings, paving the way for the engineered synthesis of complex natural products. In green chemistry manufacturing, EnzymeCAGE facilitated the design of a novel biosynthetic route to glutaric acid by accurately screening enzyme candidates for multiple consecutive reaction steps. This not only confirms the model's high reliability in constructing multi-step metabolic pathways but also provides an efficient computational framework for green alternatives to bulk chemical production.

Figure 3: Enzyme retrieval by EnzymeCAGE in the glutarate biosynthesis pathway
In summary, EnzymeCAGE establishes an integrated “structure–function–evolution” framework for enzyme–reaction matching, surpassing traditional limitations in enzyme function prediction and demonstrating strong application potential in external validations. The research team plans to further refine reaction center identification and fine-tune the model for specific enzyme families. Both the model and associated data have been made open-source, offering a reusable tool for the scientific community. It is recognized that EnzymeCAGE holds promise in shortening enzyme discovery cycles and accelerating the adoption of biocatalysis in pharmaceuticals, energy, environmental applications, and green biomanufacturing.
Paper Title:
A Geometric Foundation Model for Enzyme Retrieval with Evolutionary Insights
Paper Link:
https://www.nature.com/articles/s41929-026-01478-y
Author Profiles

Yong Liu
Doctoral student at Shanghai Jiao Tong University (Class of 2025). Research interests: AI-driven biosynthesis, enzyme discovery and design, and multi-agent system development.

Shuangjia Zheng
Tenured-track Assistant Professor and P.h.D. Supervisor of the Global Institute of Future Technology, SJTU. Prof. Zheng's research primarily focuses on the intersection of generative artificial intelligence and drug design. He has published over 60 papers in prestigious international journals and conferences, including Nat. Mach. Intell., Nat. Comput. Sci., Nat. Commun., Nat. Biomed. Eng, NeurIPS, and ICLR, with citations exceeding 5000. Several of his achievements have been reported by renowned media outlets, including MIT Tech Review, Forbes, China Science Daily, People's Daily, and Xinhua Net. He has been selected for the Asian Young Scientist Fellowship, Forbes 30 Under 30 Asia, the WAIC Yunfan Award, and the Shanghai Chenguang Program. He has also received numerous honors and awards, including the World Artificial Intelligence Conference Outstanding Paper Award, the Rey Wu Prize, the Baidu Scholarship, and the CAAI Outstanding Doctoral Dissertation Award.




